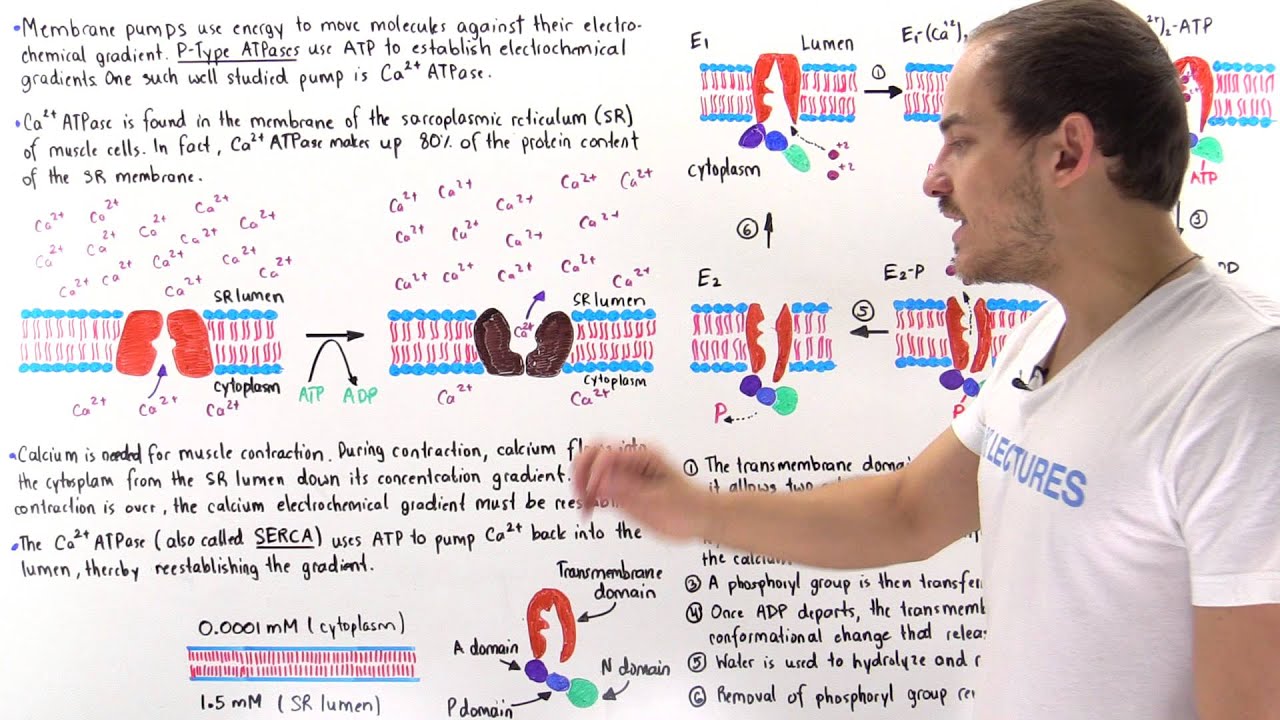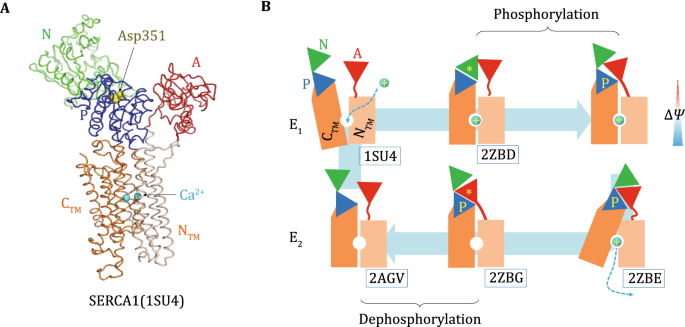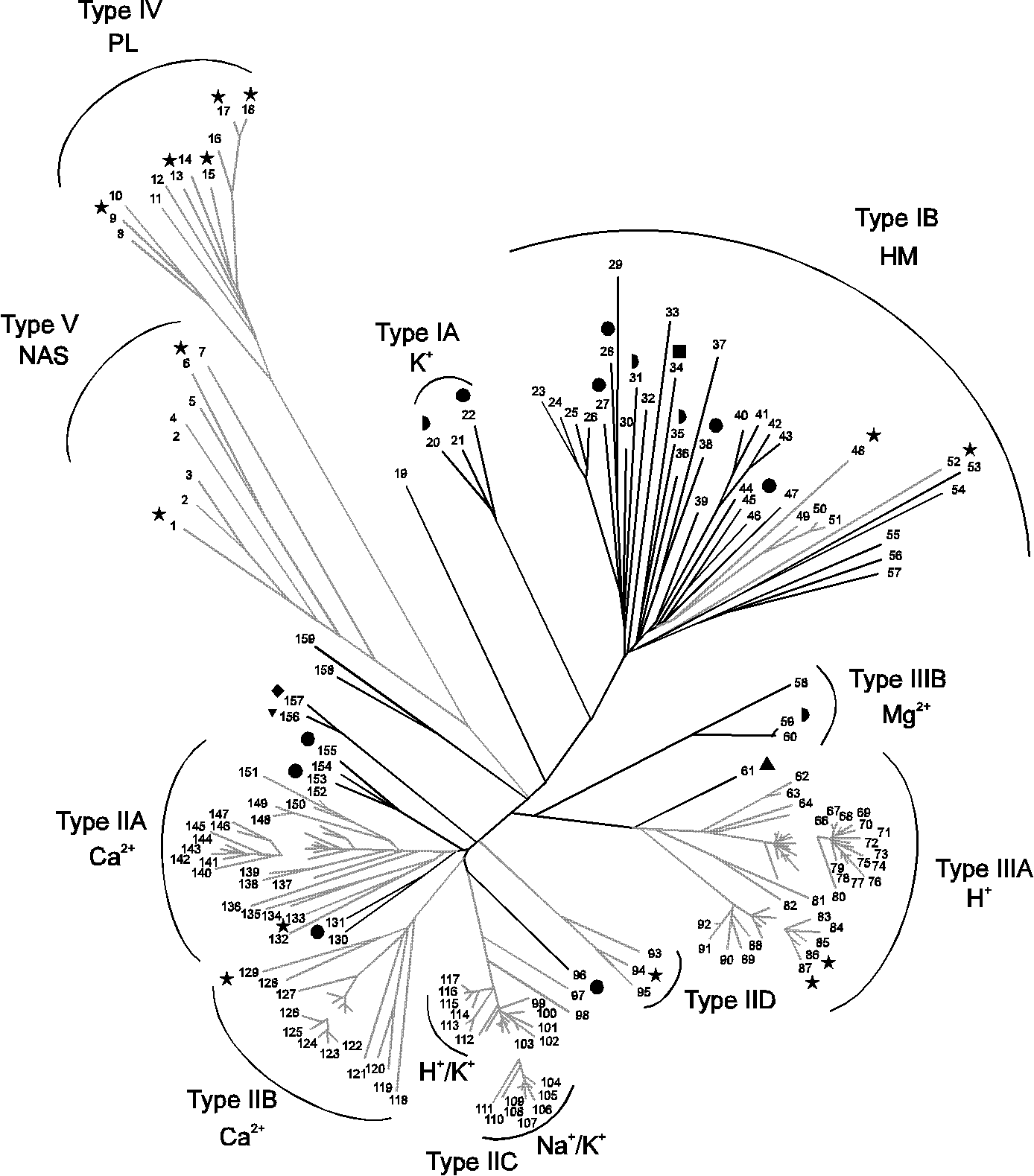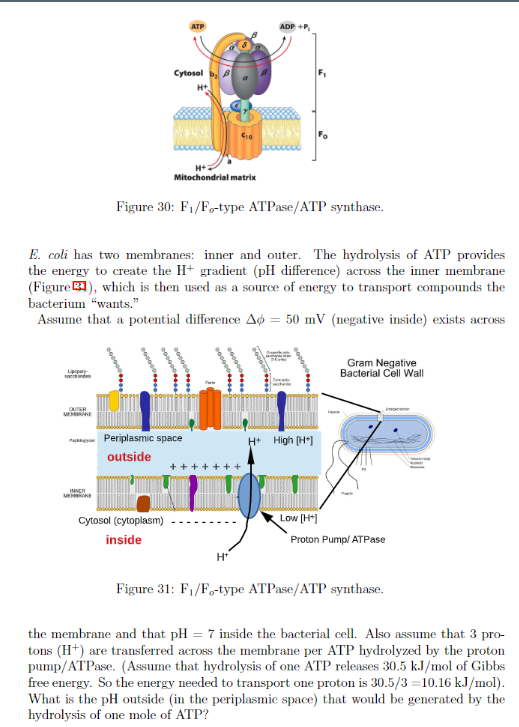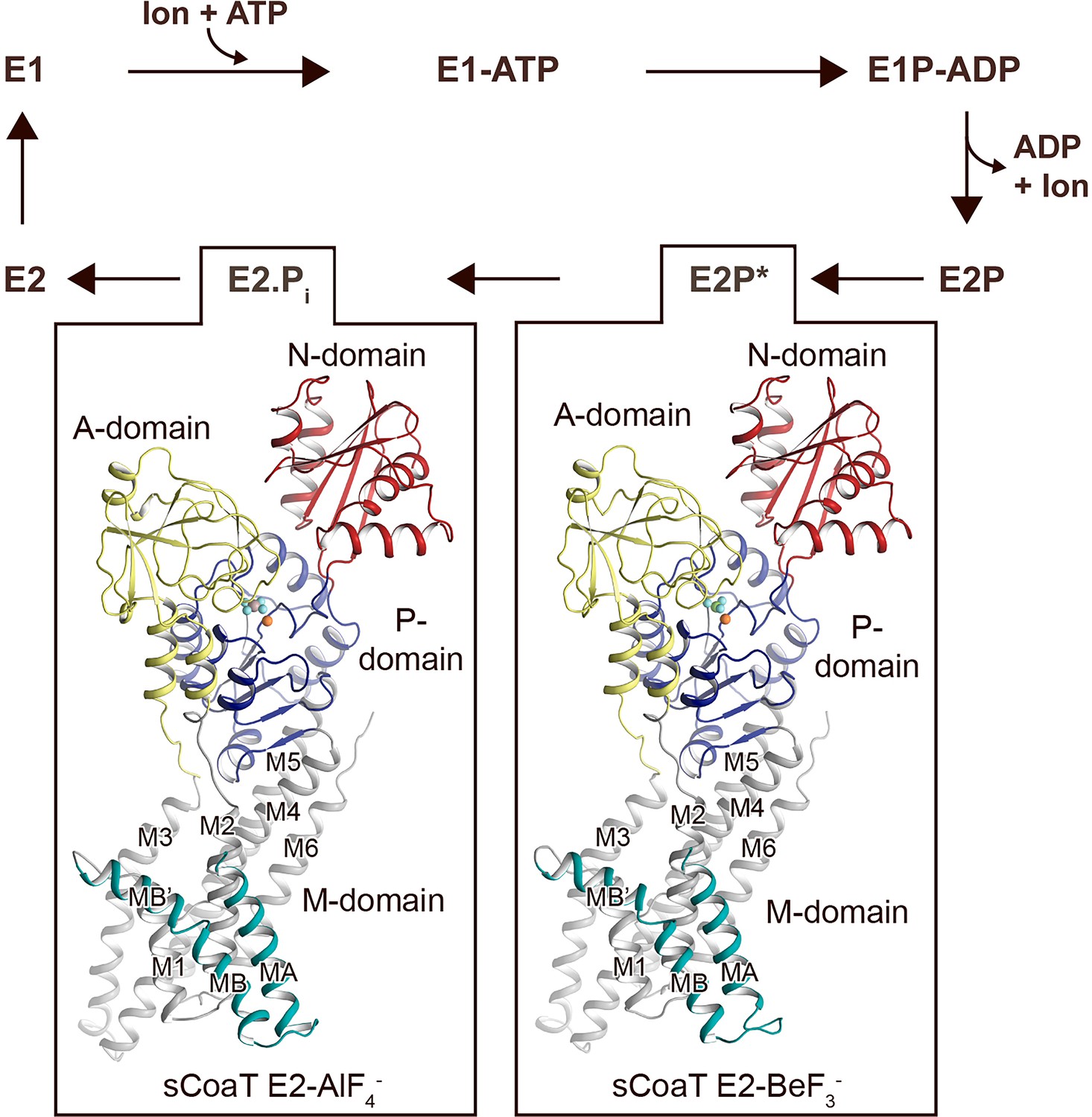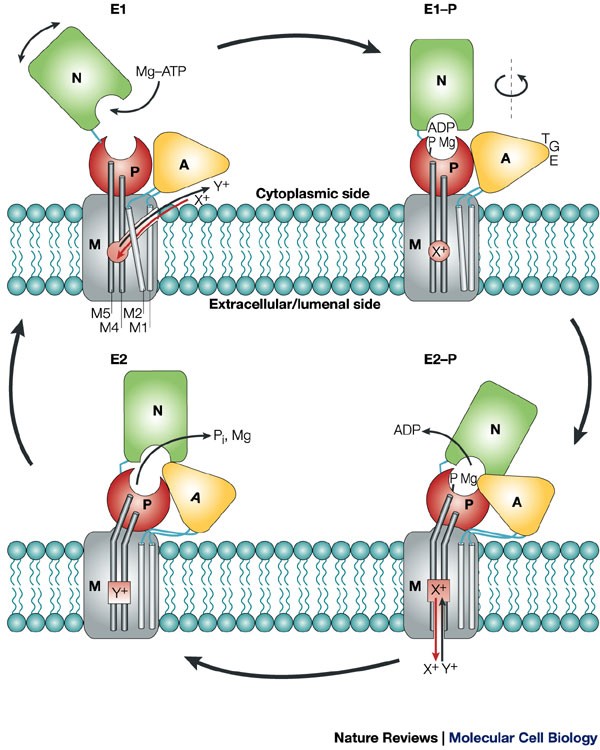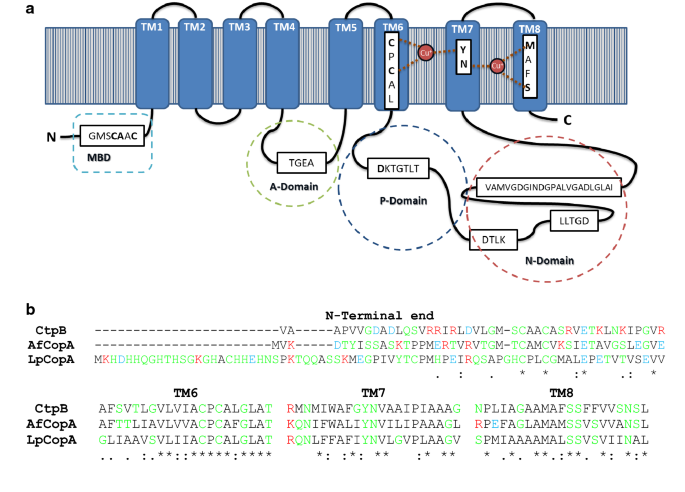
CtpB is a plasma membrane copper (I) transporting P-type ATPase of Mycobacterium tuberculosis | Biological Research | Full Text

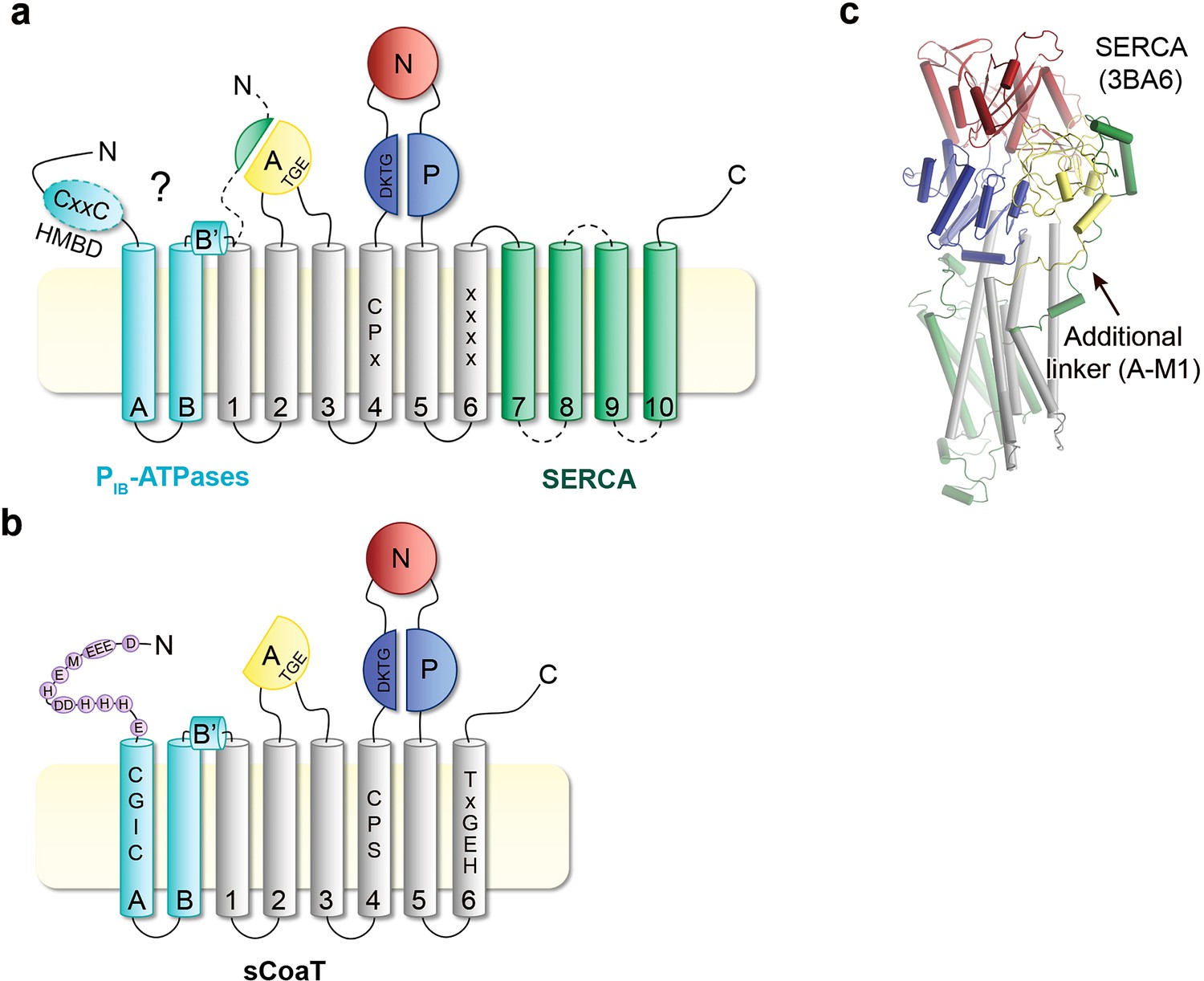
PDB structures and conservation of P-type ATPases. Diagram applicable... | Download Scientific Diagram
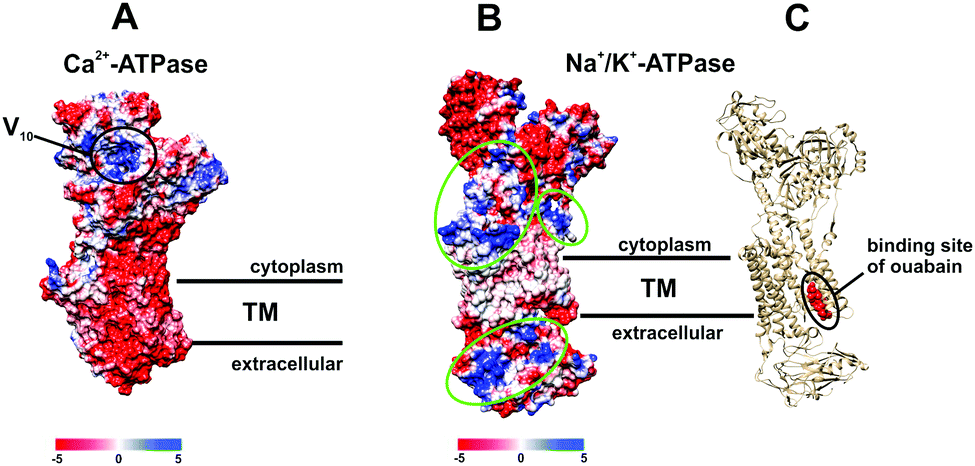
The P-type ATPase inhibiting potential of polyoxotungstates - Metallomics (RSC Publishing) DOI:10.1039/C7MT00279C

biochemistry - Difference between the P4 and P5 subtypes of P-type ATPases in plants - Biology Stack Exchange
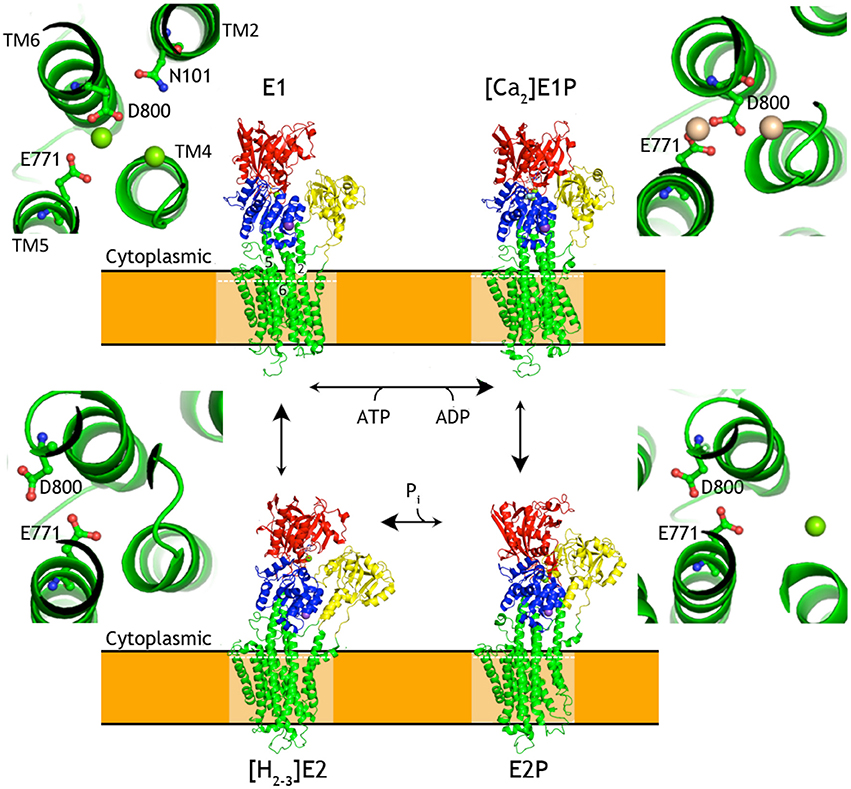
Frontiers | Improved Model of Proton Pump Crystal Structure Obtained by Interactive Molecular Dynamics Flexible Fitting Expands the Mechanistic Model for Proton Translocation in P-Type ATPases

A Novel Mechanism of P-type ATPase Autoinhibition Involving Both Termini of the Protein - Journal of Biological Chemistry

The P5A ATPase Spf1p is stimulated by phosphatidylinositol 4-phosphate and influences cellular sterol homeostasis | Molecular Biology of the Cell
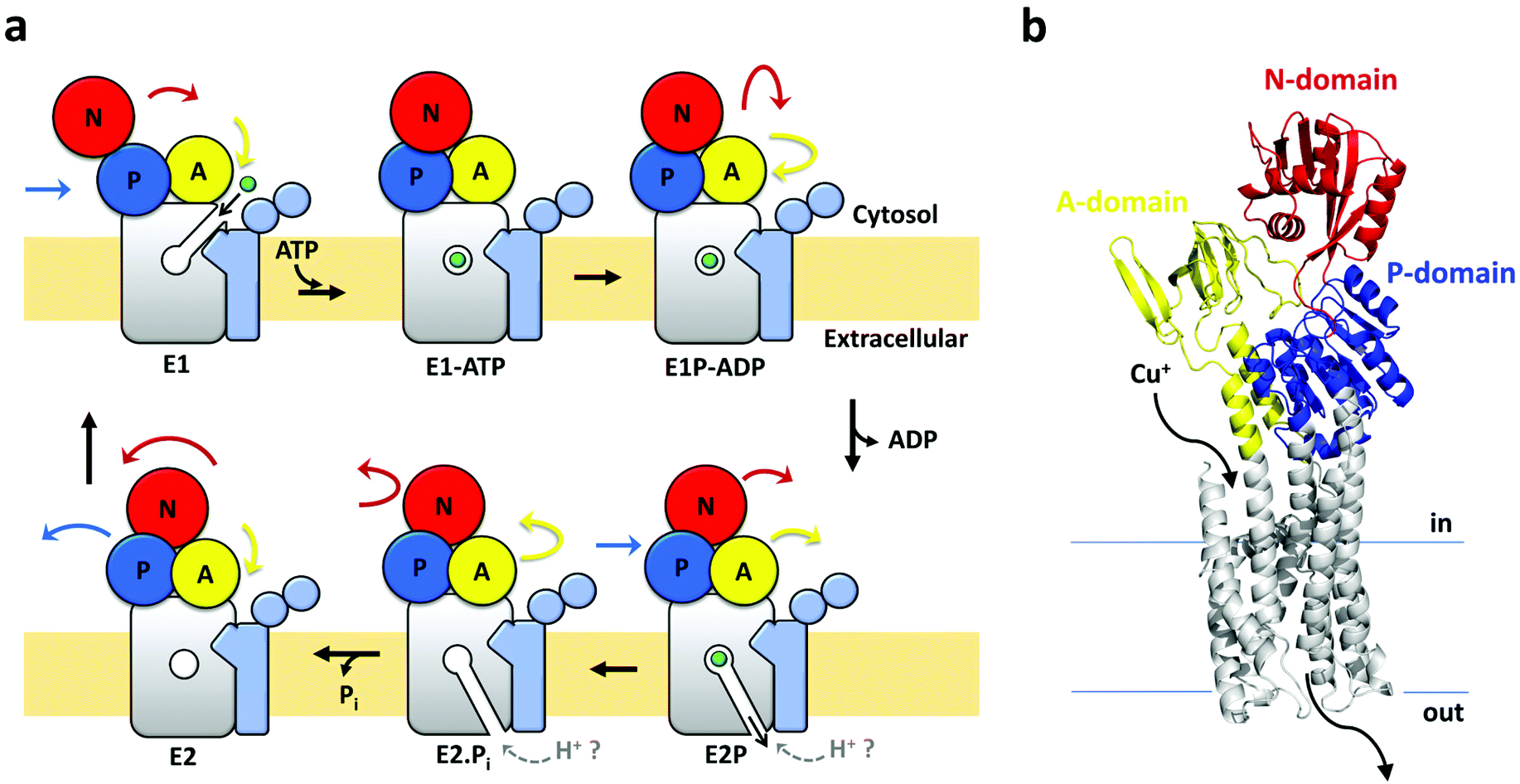
Transmembrane Cu( i ) P-type ATPase pumps are electrogenic uniporters - Dalton Transactions (RSC Publishing) DOI:10.1039/D0DT01380C

ATPase Focused Libraries | Targeted and Focused Screening Libraries | Screening Libraries | Life Chemicals

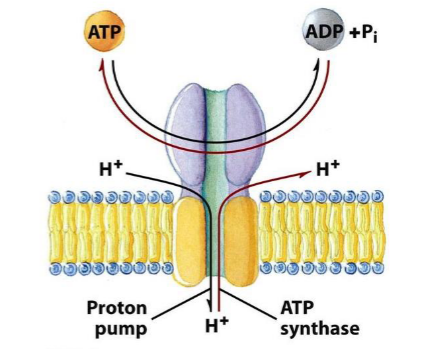
F- TYPE ATPase|| V-TYPE PUMP ||ABC-TRANSPORTER||PRIMARY ACTIVE TRANSPORT||CSIR-NET||GATE||BARC||ICMR - YouTube